Human Hexa-Histidine-Tagged Single-Chain Variable Fragments for Bioimaging of Bacterial Infections
Abstract
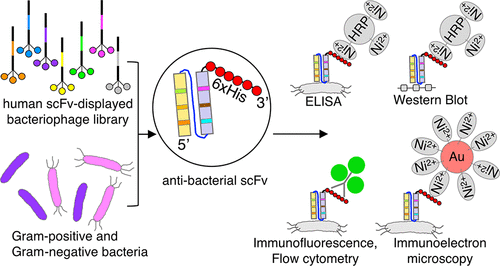
The single-chain variable fragment (scFv) of monoclonal antibodies is a promising recombinant nanostructure for various medical applications, including bioimaging and targeted therapy. While numerous scFv antibodies against eukaryotic cell surface proteins (especially cancer biomarkers) have been generated and engineered to suit various purposes, only a few specific scFv against bacterial cell surfaces have been developed, especially those of human origin. Recent incidents of emerging multidrug-resistant pathogenic bacteria and the realization of the importance of a balanced microbiota on the health of the host has led to more interests in the development of recombinant antibacterial antibodies as a detection probe or targeted therapy for bacterial infections. This study reports the generation of two specific human antibacterial scFv using phage display antibody technology. The recombinant scFv fragments of about 30 kDa and a diameter of 5 nm were produced and purified from engineered Escherichia coli that can enhance cytosolic disulfide bond formation. As a proof of principle, Propionibacterium acnes and Pseudomonas aeruginosa were used as model Gram-positive and Gram-negative bacteria, respectively. Specificity at the strain and species level to both planktonic and biofilm forms of these bacteria were demonstrated in various assay formats, namely, ELISA, flow cytometry, western blot, immunofluorescence, and electron microscopy via the hexa-histidine tag. This recombinant scFv generation platform can be applied for other bacteria, and since the scFv obtained has a benefit of being a human origin, it could be conveniently engineered for various therapeutic or theranostic applications with minimized adverse immunoreaction.
Introduction
Phage display antibody technology, one of the most popular methods for the generation of recombinant antibodies, has recently been awarded the Nobel Prize in Chemistry for the year 2018. The key advantage of this technology relies on the simplicity but powerful affinity selection (bio-panning) procedure and the direct linkage between the displayed antibody and its encoding gene within the virion genome. Recombinant antibodies against a desired target can be identified directly from diverse repertoires of antibody genes, generating high-affinity binding sites without the constraint imposed by classical methods for generating either polyclonal or monoclonal antibodies. Since this method does not depend on an animal’s immune system, antibodies to a wide variety of antigens, including molecules that cannot stimulate the immune system of animals, such as small molecule haptens and self-antigens have been successfully isolated using this technology. Various forms of antigen-binding fragments, including Fab and scFv have been cloned and displayed on the filamentous bacteriophage M13. ScFv is a nanostructure that consists of light and heavy chain variable domains of immunoglobulins of mammals, linked with a peptide linker (G4S)3. One scFv fragment has a molecular weight of about 30 kDa and a diameter of 5 nm, which is about one-fifth the size of the parental IgG molecule. Once the sequence of scFv against any target has been identified, it can be further engineered into various formats including whole immunoglobulin (IgG) monoclonal antibodies and bispecific antibodies or fabricated into various nanoparticles for both therapeutic and diagnostic purposes.Most of the research and development on recombinant antibodies mainly focuses on cancer. Several hundred monoclonal antibodies have been approved or are in clinical trial stages for the treatment of cancer and autoimmune diseases. Until now, only a handful of publications regarding the generation of recombinant antibodies against infectious micro-organisms have been published. The emergence of multidrug-resistant pathogenic bacteria as well as the concern for a balanced microbiota have led to more interest in generating recombinant antibodies against pathogenic bacteria.The model Gram-negative bacteria in this study, P. aeruginosa, is one of the WHO prioritized pathogens for which research and development of new antibiotics are critically needed. Some research groups used molecular targets, such as purified recombinant proteins of P. aeruginosa, whereas others used whole bacterial cells in suspension as targets for affinity selection. All carried out affinity selection using peptide display phage libraries, and from this, anti-P. aeruginosa peptides were identified. On the other hand, a study group from MedImmune Ltd., UK, used P. aeruginosa whole cells as a target for affinity selection against a human scFv library derived from patients who had recently recovered from P. aeruginosa infections and identified a scFv against P. aeruginosa exopolysaccharide residue (Psl). Another study group used the recombinant exotoxin A of P. aeruginosa to do affinity selection against a human scFv phage display library based on 60 Thai blood donors and isolated anti-exotoxin A scFv clones. To the best of our knowledge, only the anti-Psl scFv of MedImmune Ltd. group has been developed into an anti-pseudomonal drug candidate.The model Gram-positive bacteria of this study, P. acnes, have been used as an affinity selection target by one study group. They generated a scFv display phage library from human peripheral blood mononuclear cells, which were immunized in vitro with heat-killed P. acnes and three anti-P. acnes scFv clones were identified, but there was no known further development from those clones.Biopharmaceuticals could become a reliable resource to fight back multidrug- and pandrug-resistant bacteria, which has been a global concern. In 2017, multidrug-resistant P. aeruginosa caused an estimated 32,600 infections among hospitalized patients and 2700 estimated deaths in the US alone. It has been estimated that human deaths attributable to antimicrobial resistance (AMR) will surpass that of cancer in 2050 and the global GDP loss related to AMR will be trillions of USD. Recently, cartography of opportunistic pathogens and antibiotic resistance genes in a tertiary hospital environment, from genomic and metagenomic analysis, has been reported. Therefore, obtaining scFv nanostructures against living bacteria, which could be further engineered to be used as alternative diagnostic and therapeutic strategies to combat deadly bacterial infections, is highly attractive.This study reports an efficient method for the generation of specific human scFvs against both Gram-positive and Gram-negative bacteria using phage display technology.
 MYLAB @ SUT
MYLAB @ SUT